 The laborious process of performing and statistically analyzing genomic experiments typically culminates in a list of genes or proteins. However, deriving biological meaning from these lists is often possible only after overlaying them on known biological pathways.
The laborious process of performing and statistically analyzing genomic experiments typically culminates in a list of genes or proteins. However, deriving biological meaning from these lists is often possible only after overlaying them on known biological pathways.
A variety of bioinformatics tools are available for this task, providing sophisticated visualizations of the genes within the pathways, advanced pathway enrichment analyses, and integration of vasts amounts of knowledge from pathway databases and published literature. In addition, pathway data may be integrated with protein-protein, protein-DNA and protein-compound interactions data.
At the Bioinformatics Core Facility we have gained vast experience in downstream analysis of high-throughput genomic experiments using cutting edge pathway databases and tools such as KEGG, STRING, BioCyc, Reactome, Cytoscape, GSEA and others. In addition, we provide consultation on the use of highly popular commercial software for pathway analysis such as Ingenuity Pathway Analysis and GeneGo MetaCore. We will be happy to match the most adequate tool to any particular experiment, and to consult the user or analyze the data accordingly.
Our services include:
- Mapping IDs, names or sequences of provided molecules to corresponding entities in pathway databases
- Identifying pathways or reactions which are statistically over-represented in provided gene or protein lists
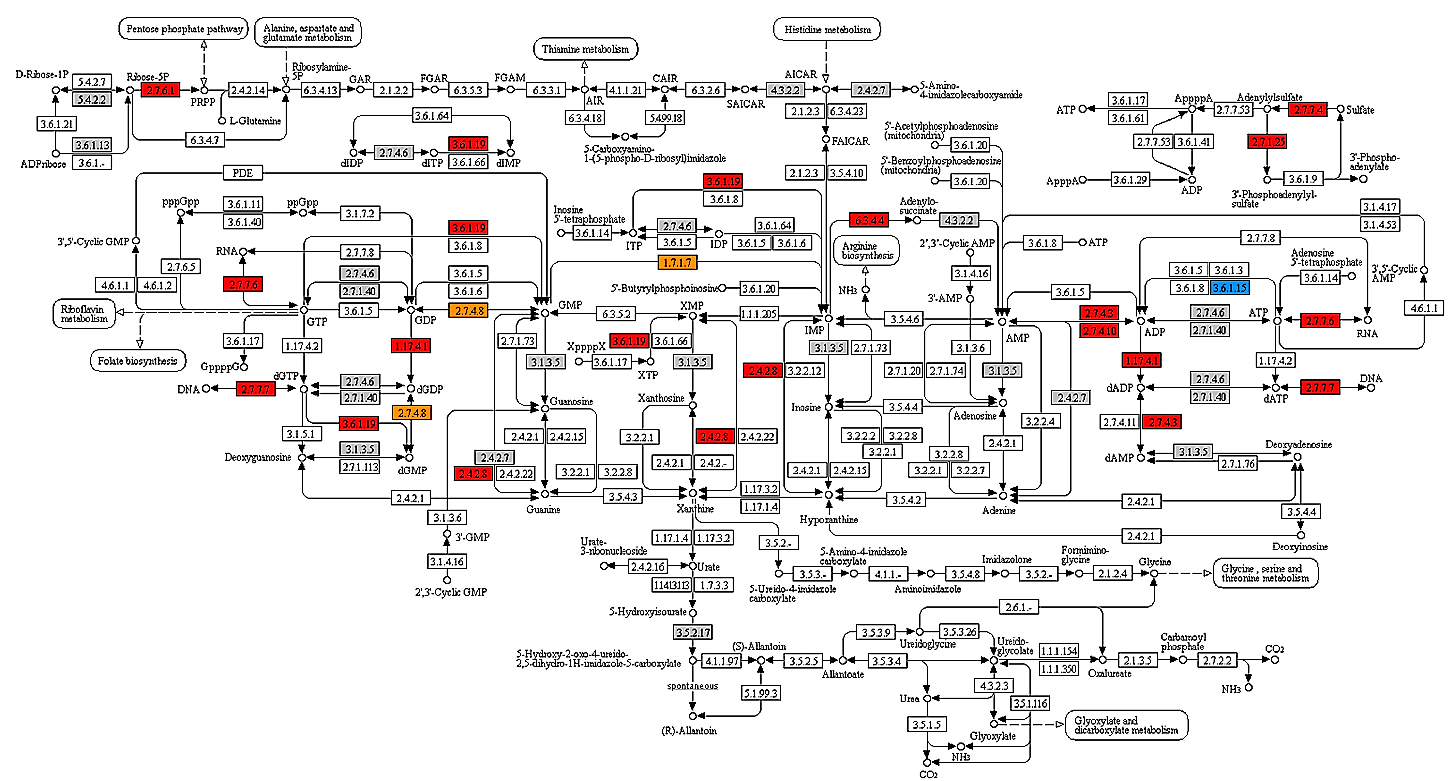
- Visually overlaying gene expression data on biological pathways and coloring the pathways according to either absolute or differential gene expression
- Visually overlaying gene expression data on protein-protein interaction networks
- View and compare equivalent pathways in various species